Infos zum Film
"Solo Sunny"
DEFA 1980, 104 min.
FSK/JMK 12 Jahre
Regie Konrad Wolf, Drehbuchautor Wolfgang Kohlhaase
Fr., 17. Feb. 2023, 18:00 Uhr - 21:00 Uhr
Im sozialistischen Alltag hatten Frauen viele Erwartungen zu erfüllen: als Werktätige, Mütter, Hausfrauen, politische Aktivistinnen. Wie ließen sich individuelle Lebenspläne und gesellschaftliche Normen vereinbaren? Darum geht es im Film und in der Diskussion der Herrenhausen Science Movie Night am 17. Februar 2023 in Hannover.
Schon 1950 war die Hälfte der Frauen in der DDR berufstätig. 1989 betrug der Anteil sogar 90 Prozent. Das war Weltspitze. Aber keineswegs ein Beleg für verwirklichte Emanzipation. Den Maßstab der Gleichberechtigung bestimmten in der DDR, genau wie im Westen: Männer. Frauen waren als Werktätige unverzichtbar zur Erreichung der Planziele. Aber sie hatten gleichzeitig Rollenerwartungen als Ehefrauen, Mütter, Hausfrauen und Aktivistinnen in gesellschaftlichen Organisationen zu erfüllen. Um die Alltagslast zu lindern, half der Staat zwar mit sozialpolitischen Maßnahmen wie kostenloser Kinderbetreuung, Haushaltstag, Frauensonderstudium. Doch Frauen, die sich nicht anpassen wollten, hatten es schwer.
Der DEFA-Film "Solo Sunny" von 1980, der in dieser Science Movie Night gezeigt wird, erzählt von einer jungen Frau, die als Band-Sängerin durch DDR-Säle tingelt und mit ihren Erwartungen ans Leben immer wieder in Konflikt gerät mit ihren Mitmenschen und dem Alltag im Sozialismus.
Nach der Filmvorführung diskutieren Expert:innen über das politische und gesellschaftliche Frauenbild in der DDR.
Dieser Film ist der zweite Film der fünfteiligen Reihe "Film trifft Wissenschaft – Das Leben in der DDR".
"Solo Sunny"
DEFA 1980, 104 min.
FSK/JMK 12 Jahre
Regie Konrad Wolf, Drehbuchautor Wolfgang Kohlhaase
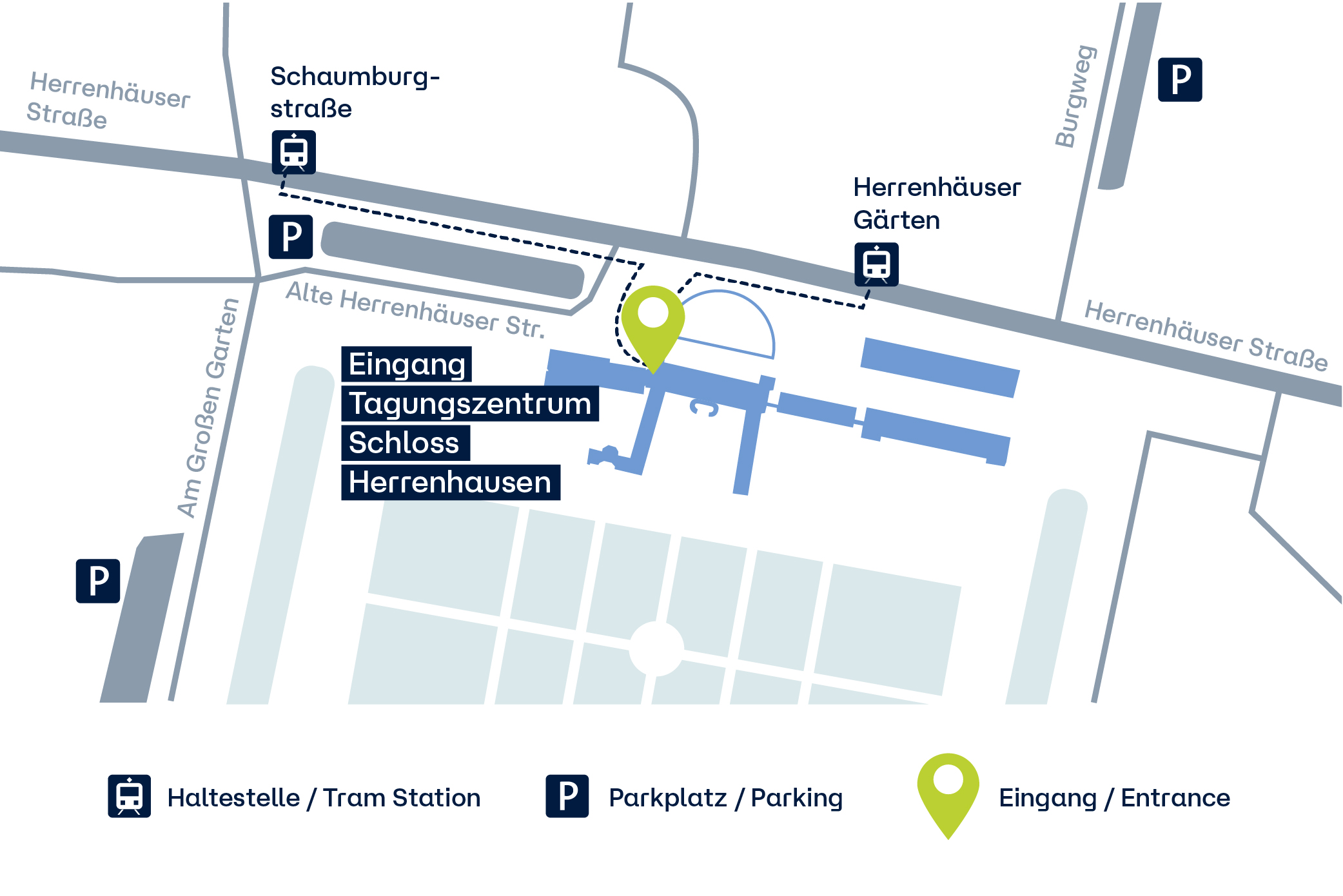
Die Veranstaltungen finden im Tagungszentrum Schloss Herrenhausen, Herrenhäuser Straße 5, 30419 Hannover statt (zu Google Maps).
Der Eingang zum Tagungszentrum (drei große Holztüren) befindet sich auf der Westseite des Schlosses, im Seitenflügel, direkt neben dem Eingang des Restaurants "Grauwinkels Schlossküche".
Schloss Herrenhausen ist barrierefrei.
Der Eintritt zu den Veranstaltungen der VolkswagenStiftung ist kostenfrei. In seltenen Fällen kann es sein, dass der Zutritt zum Tagungszentrum Schloss Herrenhausen nur mit einer (ggf. kostenpflichtigen) Eintrittskarte möglich ist. Sollte dies so sein, finden Sie einen entsprechenden Hinweis auf der Webseite der jeweiligen Veranstaltung.
Die Türen öffnen 45 Minuten vor Veranstaltungsbeginn.
Bei Veranstaltungen der VolkswagenStiftung im Schloss Herrenhausen können Sie gerne unsere kostenfreie Garderobe nutzen.
Öffentliche Abendveranstaltungen dauern ca. 100 - 120 Minuten.
Wir legen großen Wert auf ein respektvolles und wertschätzendes Miteinander. Wenn Sie mehr über diese Haltung erfahren möchten, freuen wir uns, wenn Sie auf unserer Seite "Respektvoll miteinander!" vorbeischauen.
Einige unserer Veranstaltungen werden auch im Livestream übertragen. Sie erkennen dies am Hinweis "Livestream" im Veranstaltungskalender und auf der jeweiligen Veranstaltungsseite.
Viele der Veranstaltungen werden live übertragen und aufgezeichnet. Mitschnitte und Berichte zu vergangenen Veranstaltungen finden Sie in unserem Newsroom sowie im Youtube-Kanal der Volkswagenstiftung.
Bitte kontaktieren Sie die Schloss Herrenhausen Veranstaltungs- und Betriebs GmbH unter: 0511/763 744-0.
Bei Fragen erreichen Sie unsere Hotline von 8:00 - 16:00 Uhr unter 0511/8381-200.


In der Themenreihe "Leben in der DDR" zeigen die Herrenhausen Science Movie Nights "Film trifft Wissenschaft" ausgewählte Filme, die den Alltag in der DDR illustrieren. Sie erzählen von Sehnsüchten, Hoffnungen und Träumen, die gar nicht so anders als die ihrer westdeutschen Nachbarn waren. Ganz unterschiedlich war jedoch die gelebte Realität. Wie diese aussah erkunden die fünf Filmabende mit den Schwerpunkten: Jugend, Frauen, Homosexualität und Überwachung in der DDR sowie dem Leben nach der Wende.
Hier finden Sie weitere Termine:
Fr., 10. Mär. 2023, 18:00 Uhr bis 21:00 Uhr